Table of contents
These two dataset can both be used to follow the tutorial and practice using Minerva Author. Download the files to your local computer to use them.
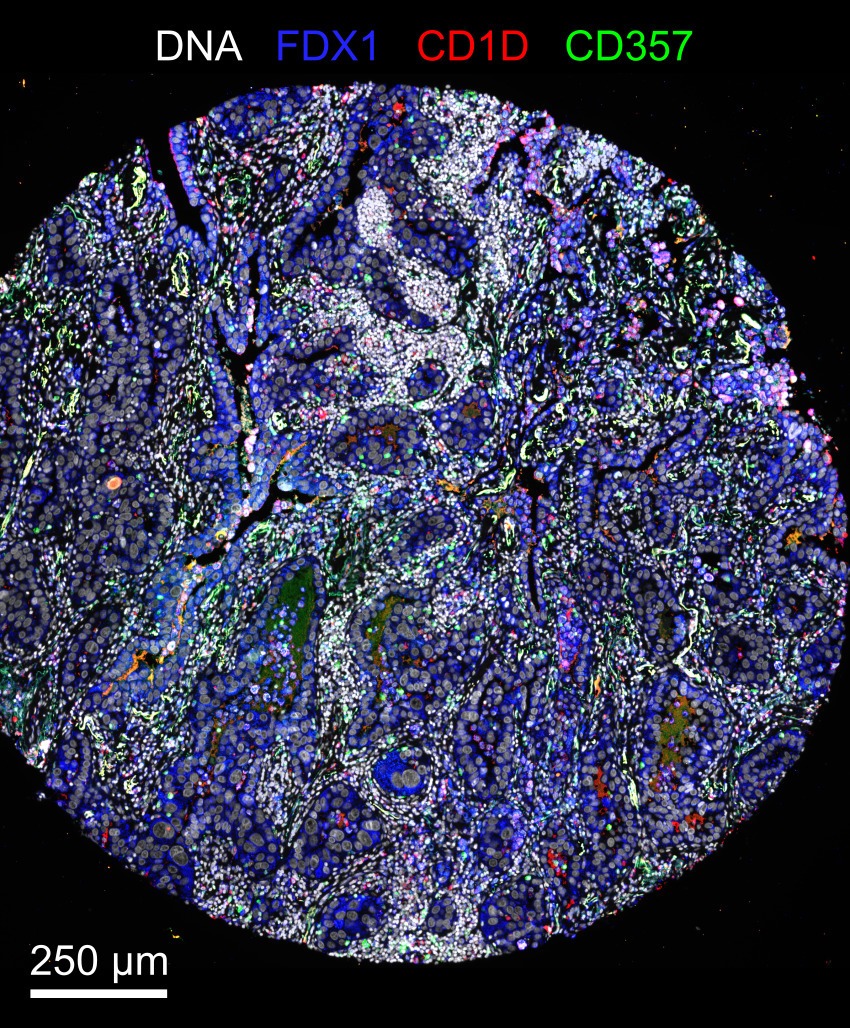
Examplar 1 - Multiplex immunofluorescnet images of small lung adenocarcinoma
Examplar 1 contains Cyclic Immunofluorescence (CyCIF) images of small lung adenocarcinoma specimens from larger tissue microarray (TMA).
Click the image below to download Examplar 1
File structure for Examplar 1
Downloading Examplar 1 will give you the following files. Below is a few words on what they are and how you can use them to practice using Minerva Author.
| File/folder name | Content |
| raw | This folder contains 3 .ome images that you can use as input for Minerva Author. |
| illumination | This folder contains .tif images that represents illumination correction for the image in raw. You do not need these for your Minerva Story. |
| markers.csv | This is .csv file contains the channel names. You can use it as input for input for Minerva Author. |
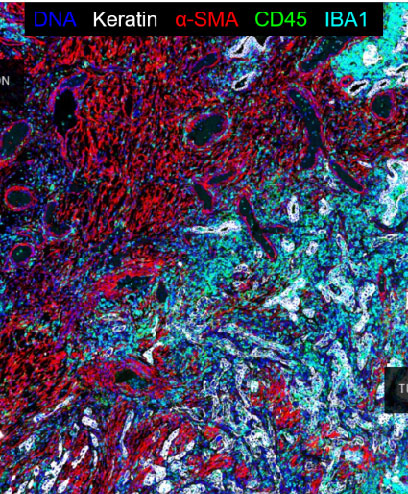
Examplar 2 - Multiplex immunofluorescent images of human lung cancer
Examplar 2 played an instrumental role in the development process of Minerva. Before using Examplar 2, keep in mind that the image file is 20GB in size. Examplar 2 also has data for visualization charts that you can use to follow the instructions to add Data Visulaization to your story.
This dataset is described in Rashid et al., Scientific Data, 2019. It comprises of multiplexed immunofluorescence images and derived single-cell measurements of immune lineage and other markers in formaldehyde-fixed and paraffin-embedded (FFPE) human lung cancer tissue. We used tissue cyclic immunofluorescence (t-CyCIF) to generate fluorescence images which we artifact corrected using the BaSiC tool, stitched and registered using the ASHLAR algorithm, and segmented using ilastik software and MATLAB. We extracted single-cell features from these images using HistoCAT software. This data is also described in Du, Lin, and Rashid et al., Nature Protocols, 2019.
For practice, you can use this dataset to try to reproduce the two stories created to describe histologic features and quantitative single-cell data analysis in a single lung cancer specimen.
Image File (20 GB)
Click the image below to view and download the image file in Examplar 2